Study identifies a new strain of zika virus circulating in Brazil
26/06/2020
Raiza Tourinho (Fiocruz Bahia)
Even in the midst of a pandemic that has affected the everyday life of virtually everyone, Brazilians are still living with the consequences of the last national public health emergency: the zika virus, which has led to the birth of 3,534 babies with congenital zika syndrome (CZS) since 2015. A new strain of the zika virus was recently discovered in Brazil by researchers of the Center for the Integration of Data and Knowledge for Health (Cidacs) of Fiocruz Bahia, and the possibility of the re-emergence of an epidemic of zika has become higher. The finding was published in early June in the International Journal of Infectious Diseases and should be taken as a warning for surveillance of the disease.
According to the latest epidemiological report by the Ministry of Health, of the main arbovirus diseases circulating in Brazil, zika has been the one with the lowest number of cases in 2020: 3,692 probable cases were reported (incidence rate of 1.8 cases per 100,000 inhabitantes), as opposed to the 47,105 probable chikungunya cases (incidence rate of 22.4 cases per 100,000 inhabitantes) and 823,738 probable dengue fever cases (incidence rate of 392 cases per 100,000 inhabitantes). But this situation may change if a new genetic strain begins to circulate in the population.
Tool
The introduction of a new lineage in the country was identified by a genetic monitoring tool developed by researchers linked to Cidacs and to the Gonçalo Moniz Institute (Fiocruz Bahia); Faculty of Technology and Sciences (FTC); University of Salvador (Unifacs), and Escola Bahiana de Medicina e Saúde Pública (Bahia School of Medicine and Public Health - EBMSP). The researcher of Cidacs Biotechnology Information Platform, Artur Queiroz, one of the leaders of the study, explains that the tool developed by the group analyzes sequences available in public data bases and makes it possible to identify the zika virus strains present in data bases of the National Center for Biotechnology Information (NCBI).
“We collect these data and analyze them, then select the Brazilian sequences and show the frequency of these viral types year by year. The main finding is that we have seen a variation of subtypes and strains over the years, and in 2019 we detected the appearance, albeit small, of a strain that had not yet been described in circulation in the country”, he explains.
Identification
Two lineages of the zika virus are known: the Asian strain and the African strain, the latter subdivided in eastern and western). The tool analyzed 248 Brazilian sequences submitted to databases since 2015. Up to 2018, the genetic data found were mostly Cambodjan (more than 90%), a ratio that went through a radical change in 2019, when the Micronesia strain took over and was responsible for 89.2% of the sequences submitted to the genetic database.
But the thing that surprised the researchers was the identification of the emerging African type, until then unheard-of in Brazil. “The African lineage was isolated in two different regions in the country: in the South, coming from the southernmost state of Rio Grande do Sul, and in the South-East region, more specifically in Rio de Janeiro”, the study informs.
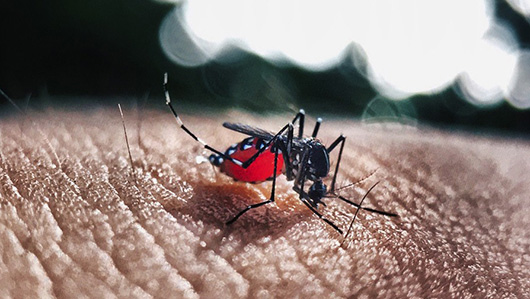
The geographical distance and the different hosts (one was found in a mosquito of the same genus of Aedis aegypti, the Aedes albopictus, and the other in a monkey species) suggest that this lineage has been circulating in the country for a while, and may have epidemic potential, once most of the population has no antibodies for this new lineage of virus. For Queiroz, the finding shows the usefulness of the tool as a “good mechanism for surveillance and alert for the possibility of a new zika virus epidemic”.
“With most of everyone’s attention focused on Covid-19 these days, this study is a warning to keep us from forgetting other diseases, zika in particular. Due to the circulation of the virus in the country, new genetic studies must continue to be carried out to avoid a new outbreak of the disease with the new genotype”, stresses Larissa Catharina Costa, one of the authors of the study. The tool developed by the researchers can be found here. The study itself can be found here.